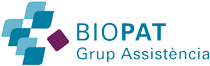Montse Verdú1,2, Ruth Román1, Miquel Calvo3, Natàlia Rodón1, Beatriz García1, Marta González1, August Vidal1,2, and Xavier Puig1,2,4.
1BIOPAT. Biopatologia Molecular, SL, Grup Assistencia, Barcelona, Spain; 2 Histopat Laboratoris, Barcelona, Spain; 3Statistics Department, Universitat de Barcelona, Barcelona, Spain and 4Hospital de Barcelona, SCIAS, Grup Assistencia, Barcelona, Spain.
Abstract
Invasive micropapillary carcinoma is associated with frequent lymph node metastasis and adverse clinical outcome. Initially described as a variant of breast and ovarian carcinoma, it has subsequently been found in other organs, most recently the colon. Reports of colorectal micropapillary carcinoma to date are limited in number, and their molecular profile has not been established. The aims of the present study were to analyze their clinicopathological features and molecular profile and compare them with those of conventional adenocarcinoma. Clinicopathological features of a cohort of 379 patients with primary colorectal cancer were retrospectively reviewed for the presence of the pattern characteristic of micropapillary carcinoma. We also assessed the expression of KRT7, KRT20, CEACAM5, MUC1 (EMA, clone E29), MUC1 (clone MA695), MLH1, MSH2, MSH6 and TP53 by immunohistochemistry. Genetic assessments of microsatellite instability, chromosomes 17p and 18q, and mutations in TP53, BRAF and KRAS were performed using DNA extracted from formalin-fixed, paraffin-embedded sections. In all, 60 of the reviewed cases (16%) had a micropapillary component that ranged from 5 to 95% of the tumor, characterized by a higher frequency of an infiltrative pattern, lymphovascular and perineural invasion, a higher depth of invasion and more positive lymph nodes than conventional adenocarcinoma. Immunohistochemistry for MUC1 (clone MA695) and MUC1 (EMA, clone E29) enhanced the characteristic inside-out staining pattern of the micropapillary carcinoma component, whereas the rest of the tumor showed luminal staining patterns. KRT7 expression was slightly increased in micropapillary carcinoma, but did not reach significance (17-3%, p=0.1967). The molecular parameters showed a higher frequency of TP53 alterations and a low incidence of microsatellite instability and RER phenotype (loss of mismatch repair protein) in micropapillary carcinoma. With regard to the histological parameters, micropapillary carcinoma appears to be more aggressive than conventional colorectal adenocarcinoma. The molecular profile supports the hypothesis that micropapillary carcinoma carcinogenesis develops through the classical chromosomal instability pathway.
Keywords: clinicopathological characterization; colorectal cancer; immunohistochemistry; micropapillary carcinoma; molecular characterization; MUC1.
Introduction
Micropapillary carcinoma is accepted as an aggressive variant of colorectal adenocarcinoma, characterized histologically by small papillary cell clusters surrounded by lacunar spaces. It was described first in the breast and ovary, and subsequently in other organs, including the bladder, lung, pancreas and salivary glands.1-7
Previously published work found that the proportion of micropapillary carcinoma to the entire tumor ranged from 5 to 80% but was usually less than 30% of the entire lesion, with no pure cases of micropapillary carcinoma reported. Most cases were located in the sigmoid colon and the rectum. 8-13
Other studies have shown that micropapillary carcinoma is more commonly associated with lymphovascular invasion, lymph node metastasis and aggressive biological and clinical behavior. The inverted polarity of the cells that compose the micropapillary nests with an “inside-out” growth pattern, which has been shown by immunohistochemical and ultrastructural studies14,15 is probably related to its high invasive potential.
Fewer than 130 cases of colorectal micropapillary carcinoma have been reported. The aim of the present study was to define the molecular profile of this variant, which has thus far not been well characterized because of a lack of clinical data.8-13
Material and methods
Patients, tissues and clinicopathological features
A cohort of 379 cases of primary colorectal adenocarcinoma was retrieved from the archival files of Histopat Laboratories (Barcelona, Spain); cases were consecutive and collected between 1993 and 2008. Hematoxylin and eosin-stained sections (average four slides per tumor) were retrospectively reviewed by two pathologists to detect, quantify and localize a distinct micropapillary carcinoma pattern. We now characterize micropapillary carcinoma as neoplastic cell clusters with papillary morphology but without a true fibrovascular core, surrounded by empty lacunar spaces lined by strands of fibrocollagenous stroma. The tumor cells show eosinophilic cytoplasm, pleomorphic, hyperchromatic nuclei and a peculiar “inside-out” pattern of cell arrangement. 1-7
Clinicopathological features, summarized in Table 1, include the following parameters: gender and age of the patient, tumor location (proximal or distal to the splenic flexure), macroscopic type (ulcerated/stenosing or polypoid/exophytic), microscopic type (conventional adenocarcinomas or mucinous, when more than 50% of the tumor has an extra- or intracellular mucoid component)16 and histological grade defined by the ratio of glandular to solid pattern, following the WHO classification16 and categorizing mucinous adenocarcinoma as grade 3. The extent of tumor invasion and lymph node involvement was assigned according to the TNM system (sixth edition)17 and the presence or absence of metastasis and peritoneal invasion is reported. Clinical stage was classified based on Astler-Coller’s system.17 To classify the growth pattern, tumors were labeled as infiltrative when irregular penetrating or permeative growth was demonstrated on the periphery of the tumor, and as expansive when the tumor border was a smooth-pushing front.18 Cytological grading was divided into three categories: high, intermediate, and low grade. Intramural and extramural thin-walled vessel invasion, venous vessel invasion and perineural invasion were also evaluated and recorded following the WHO classification.16 Thin-walled vessels are defined as vessels with only an endothelial cell layer in their walls, as opposed to venous vessels that have a muscular wall. The presence of a Crohn-like lymphoid reaction was reported when at least three nodular aggregates of lymphocytes deep to the advancing margin of the tumor were found within a single low-magnification field (4x). The presence of tumor-infiltrating lymphocytes was characterized by four or more unequivocal intraepithelial lymphocytes in a single high-magnification field (40x) of an H&E-stained section, identified and counted in areas displaying the highest content of tumor-infiltrating lymphocites.19 The tumor size was reported as the diameter maximum in millimeters. The distribution of different tumor patterns, including solid and mucinous, was reported in each case. The percentage of cribriform structures, consisting of a growing pattern of glands exhibiting small-sized secondary lumina and rounded in shape, was scored as an independent feature. Finally, the total number of lymph nodes detected and dissected was detailed.
The percentage of micropapillary component was recorded, and the case was classified as micropapillary carcinoma when it composed at least 5% of the tumor volume.
Table 1 Fisher’s exact test/Wilcoxon signed-rank test of clinicopathological features and micropapillary component
____________________________________________________________________________________________________________________________
| Categorical variable | % MC≥5 (n=60) | % MC<5 (n=319) | P-value |
| Gender | 0.6690 | ||
| Male | 37 (61.7) | 185 (58.0) | |
| Female | 23 (38.3) | 134 (42.0) | |
| Location | 0.6621 | ||
| Proximal | 24 (40.0) | 133 (35.4) | |
| Distal | 36 (60.0) | 196 (61.4) | |
| Unknown | 0 | 10 (3.1) | |
| Configuration | 0.0236 | ||
| Ulcerated/stenosing | 42 (70.0) | 171 (53.6) | |
| Polipoyd/exophitic | 18 (30.0) | 145 (45.5) | |
| Unknown | 0 | 3 (0.9) | |
| Histology | 0.0829 | ||
| Adenocarcinoma | 57 (95.0) | 275 (86.2) | |
| Mucinous | 3 (5.0) | 43 (13.5) | |
| Grade (WHO) | 0.0018 | ||
| 1 | 23 (38.3) | 171 (53.6) | |
| 2 | 30 (50.0) |
84. 26.3
|
|
| 3 | 7 (11.7) | 64 (20.1) | |
| 4 | 0 | 0 | |
| Extent of invasion (pT) | 0.0009 | ||
| pT1 | 3 (5.0) | 18 (5.6) | |
| pT2 | 2 (3.3) | 51 (16.0) | |
| pT3 | 23 (38.3) | 157 (49.2) | |
| pT4 | 32 (53.3) | 93 (29.2) | |
| Stage | 6.29 x 10-7 | ||
| 1 | 3 (5.0) | 59 (18.5) | |
| 2 | 9 (15.0) | 127 (39.8) | |
| 3 | 38 (63.3) | 108 (33.9) | |
| 4 | 10 (16.7) | 25 (7.8) | |
| Lymph node involvement (pN) | 8.49 x 10 -9 | ||
| 0 | 12 (20.0) | 192 (60.2) | |
| 1 | 22 (36.7) | 74 (23.2) | |
| 2 | 26 (43.3) | 53 (16.6) | |
| Growth pattern | 6.92 x 10-9 | ||
| Infiltrative | 58 (96.7) | 199 (62.4) | |
| Expansive | 2 (3.3) | 118 (37.0) | |
| Unknown | 0 | 2 (0,6) | |
| Cytological grading | 0.0031 | ||
| High grade | 40 (67.0) | 145 (45.5) | |
| Low or intermediate grade | 20 (33.0) | 172 (53.9) | |
| Unknown | 0 | 1 (0.3) | |
| Peritoneal invasion | 0.0012 | ||
| Present | 30 (50.0) | 92 (27.6) | |
| Absent | 30 (50.0) | 231 (72.4) | |
| Metastasis (pM) | 0.0479 | ||
| Present | 10 (16.7) | 25 (7.8) | |
| Absent | 50 (83.3) | 294 (92.2) | |
| Thin-walled vessel invasion | 7.47 x 10-12 | ||
| Present | 51 (85.5) | 120 (37.6) | |
| Absent | 9 (15) | 198 (62.1) | |
| Unknown | 0 | 1 (0.3) | |
| Venous vessel invasion | 1.86 x 10-5 | ||
| Present | 28 (46.7) | 61 (19.1) | |
| Absent | 32 (53.3) | 258 (80.9) | |
| Perineural invasion | 0.0006 | ||
| Present | 18 (30.0) | 37 (11.6) | |
| Absent | 42 (70.0) | 282 (88.4) | |
| Crohn-like lymphoid reaction | 0.0571 | ||
| Present | 15 (25.0) | 122 (38.2) | |
| Absent | 45 (75.0) | 197 (61.8) | |
| TIL | 0.5350 | ||
| Present | 6 (10.0) | 44 (13.8) | |
| Absent | 54 (90.0) | 274 (85.9) | |
| Unknown | 0 | 1 (0.3) | |
| Adenomas | 0.4169 | ||
| Present | 18 (30.0) | 78 (24.5) | |
| Absent | 41 (68.3) | 236 (74.0) | |
| Unknwon | 1 (1.7) | 5 (1.6) | |
____________________________________________________________________________________________________________________________
| Numerical variable | % MC ≥5 (n=60) | Median | % MC <5 (n=319) | Median | P-value |
| Mean ± s.d. | (IQ range) | Mean ± s.d. | (IQ range) | ||
| Age (years) | 65.8±11.7 | 66.5 (16.0) | 69.3±12.0 | 71 (15.5) | 0.0177 |
| Tumor size (max. Ø,mm) | 37.5±12.3 | 49 (15.0) | 43.8±21.0 | 40 (25.0) | 0.0429 |
| Solid carcinoma (%) | 11.5±15.2 | 10 (16.25) | 8.5 ±19.2 | 0 (10.0) | 0.0003 |
| Mucinous carcinoma (%) | 7.3±15.3 | 0 (10.0) | 13.1±(26.4) | 0 (10.0) | 0.6345 |
| Cribiform structures (%) | 9.3±18.5 | 0 (10.0) | 8.1±15.7 | 0 (10.0) | 0.7652 |
| Nodal involvement (n) | 3.5±3.3 | 2.5 (5.0) | 1.9±3.9 | 0 (2.0) | 2.76 x 10-8 |
Abbreviations: MC, micropapillary component; TILs, tumor-infiltranting lymphocites.
Bold values are statistical significance.
Immunohistochemistry and molecular studies
The immunohistochemistry profile included the expression of KRT7 (CK7), KRT20 (CK20), CEACAM5 (CEA), MUC1 (EMA, clone E29), MUC1 (clone Ma695) and TP53 and mismatch Repair gene (MMR) expression. Molecular studies performed were TP53 mutation, 17p and 18q losses, microsatellite instability status, KRAS mutation and BRAF V600E mutation; all summarized in Table 2.
Table 2 Fisher’s exact/Wilcoxon signed-rank test of immunohistochemistry expression, molecular features and micropapillary component
____________________________________________________________________________________________________________________________
| IHC expression | % MC ≥5% | % MC <5% | P-value |
| (n=30) | (n=30) | ||
| KRT7 | 0.1967 | ||
| Positive | 5 (16.7) | 1 (3.3) | |
| Negative | 225 (83.3) | 29 (96.7) | |
| KRT20 | 1 | ||
| Positive | 29 (96.7) | 28 (93.3) | |
| Negative | 1 (3.3) | 2 (6.7) | |
| 1 | |||
| CEACAM5.2 | 1 | ||
| Positive | 30 (100) | 30 (100) | |
| Negative | 0 (0) | 0(0) |
____________________________________________________________
| Molecular variable | % MC ≥5% | % MC <5% | P-value |
| (n=60) | (n=319) | ||
| TP53 accumulation | 0.1191 | ||
| Present | 39 (65.0) | 169 (53.0) | |
| Absent | 21 (35) | 149 (46.7) | |
| Unknown | 0 | 1 (0.3) | |
| TP53 mutation | 0.0564 | ||
| Present | 45 (75.0) | 193 (60.5) | |
| Absent | 15 (25) | 121 (37.9) | |
| Unknown | 0 | 5 (1.6) | |
| TP53 altered | 0.0343 | ||
| (mutation and/or accumulation) | |||
| Present | 48 (80.0) | 208 (65.2) | |
| Absent | 12 (20.0) | 108 (33.9) | |
| Unknown | 0 | 3 (0.9) | |
| 17p loss | 0.4478 | ||
| Present | 27 (45.0) | 141 (44.2) | |
| Absent | 26 (43.3) | 106 (33.2) | |
| Unknown | 7 (1.2) | 72 (22.6) | |
| 18q loss | 0.5409 | ||
| Present | 41 (68.3) | 179 (56.1) | |
| Absent | 17 (28.3) | 93 (29.2) | |
| Unknown | 2 (3.3) | 47 (14.7) | |
| MSI (NCI panel) | 0.0236 | ||
| MSS | 51 (85.0) | 240 (75.2) | |
| MSI-L | 3 (5.0) | 21 (6.6) | |
| MSI-H | 1 (1.7) | 39 (12.2) | |
| Unknown | 5 (8.3) | 19 (6.0) | |
| MSI | 0.0123 | ||
| (NCI + 17p and 18q panel) | |||
| MSS | 49 (81.7) | 215 (67.4) | |
| MSI-L | 4 (6.7) | 47 (14.7) | |
| MSI-H | 1 (1.6) | 34 (10.7) | |
| Unknown | 6 (10.0) | 23 (7.2) | |
| MMR loss of expression | 0.0629 | ||
| Present | 1 (1.7) | 27 (8.5) | |
| Absent | 59 (98.3) | 282 (88.4) | |
| Unknown | 0 | 10 (3.1) | |
| RER phenotype | 0.0105 | ||
| (MMR loss and/or MIN) | |||
| Present | 1 (1.7) | 40 (12.5) | |
| Absent | 54 (90.0) | 251 (78.7) | |
| Unknown | 5 (8.3) | 28 (8.8) | |
| KRAS mutation | 0.4641 | ||
| Present | 27 (45.0) | 121 (37.9) | |
| Absent | 30 (50.0) | 172 (53.9) | |
| Unknown | 3 (5.0) | 26 (8.2) | |
| BRAF V600E mutation | 0.1429 | ||
| Present | 9 (15.0) | 26 (8.2) | |
| Absent | 46 (76.7) | 257 (80.6) | |
| Unknown | 5 (8.3) | 36 (11.3) |
Abbreviations: MC, micropapillary component; MMR, mismatch repair; MSI, microsatellite instability; MSI-H, high microsatellite instability; MSI-L, low microsatellite instability; MSS, microsatellite stable.
Bold values are statistical significance.
Immunohistochemical analyses
All immunohistochemistry analyses were performed following an avidin-biotin immuno-peroxidase procedure. Immunohistochemistry was performed on 4-μm-thick paraffin sections after rehydration. Endogenous peroxidase activity was inhibited with 0.1% hydrogen peroxide; deparaffinized sections were pretreated to induce epitope retrieval and were independently incubated with their corresponding monoclonal antibody. Secondary reagents were biotinylated horse anti-mouse antibodies (Vector Laboratories Inc., Burlingame, CA, USA) and avidin-biotin complexes (Vectastain ABC PK 4000 ST, Vector Laboratories). 3,3′-diaminobenzidine was used as the final chromogen and Harris modified hematoxylin as the nuclear counterstain. Positive and negative controls—in which the primary antibody was omitted—were included in each experiment.
Immunohistochemistry studies for KRT7, KRT20, CEACAM5, MUC1 (EMA, clone E29) and MUC1 (clone Ma695) were performed in a subset of 60 cases. A total of 30 consecutive cases of conventional adenocarcinoma, out of which 11 were located on proximal colon (37%) and 19 on distal colon (63%); and 30 cases of micropapillary carcinoma corresponding to the cases with more proportion of this component (≥10%), out of which 14 were located on proximal colon (47%) and 16 on distal colon (53%). These analysis were carried out using primary mouse monoclonal antibodies, clones OV-TL12/30, Ks20.8, II-7 and E29 (Dako) and Ma695 (Novocastra), respectively. Immunohistochemistry evaluation was conducted using a double-blind method of scoring the presence or absence of expression in the tumor cells. Any proportion of positivity was considered.
MMR gene expression was assessed using the following primary monoclonal antibodies: anti-hMLH1 (clone G168-15), anti-hMSH2 (clone G219-1129) and anti-hMSH6 (clone 44) (BD Biosciences Pharmingen). In the tumor tissue, the absence of nuclear staining together with a positive internal control (normal mucosa or stromal cells, endothelial cells, lymphocytes) was considered as negative. Specimens with positive staining in a small cluster of neoplastic cells were considered to express the proteins. 20
Detection of TP53 protein was carried out using primary mouse monoclonal clone DO-7 (Dako). Immunohistochemistry semi-quantitative double-blind evaluation was conducted by scoring the estimated percentage of tumor cells showing nuclear staining.
Tissue macrodissection and DNA isolation
Ten 5-μm-thick sections of formalin-fixed, paraffin-embedded tissues were used for each paired case (normal and tumoral samples) to perform manual scraping. DNA was isolated using a proteinase K-phenol/chloroform protocol.21 In each specific PCR reaction, 200 ng of DNA was used, following assessment of DNA quality by PCR amplification of a 268-bp fragment of the HBB (human ß-globin) gene,22 using a GeneAmp PCR System 9700 thermal cycler (PE Biosystems, Foster City, CA, USA).
Mutational analyses
Amplicons 224 and 107 bp in size were obtained from single PCR reactions that spanned exon 15 of the BRAF gene and exon 1 of the KRAS gene. Direct sequencing of the purified amplicons was performed using the ABI PRISM® BigDye® Terminator v1.1 Cycle Sequencing kit (Applied Biosystems, Warrington, UK). Exons 4-8 of the TP53 gene were amplified in five independent PCR reactions. Mutational analysis was performed by SSCP in polyacrylamide non-denaturing pre-cast gels with two different electrophoresis conditions. Samples showing anomalous mobility were reamplified and their mutation confirmed by bidirectional sequencing.
Detection of allelic losses
Amplification of the TP53 CA dinucleotide repeat was performed for the analysis of allelic losses at chromosome 17p (TP53 locus). Loss of heterozygosity (LOH) at the chromosome 18q region involving the DCC gene was analyzed using five microsatellite markers (D18S55, D18S58, D18S61, D18S64 and D18S69) by multiplex PCR. After PCR amplification, the fluorescent products were separated by capillary electrophoresis and analyzed using the GeneScan or GeneMapper software (PE Applied Biosystems). To calculate the LOH, peak heights of both alleles from normal and tumor samples were measured in relative fluorescent units. A LOH event was considered when the ratio (N2xT1)/(N1xT2) was less than 0.5 or higher than 2.0. LOH was evaluated for each marker, and chromosome 18q allelic loss was considered when LOH was detected in at least one marker. Unstable and non-informative results for each marker were also recorded.
Analysis of microsatellite instability
Microsatellite instability status was evaluated using the five microsatellites from the NCI panel (BAT25, BAT26, D5S346, D2S123 and D17S250) in a multiplex PCR reaction. 23-25 Fluorescent amplicons were analyzed on an automated ABI PRISM–310 Genetic Analyzer using GeneScan or GeneMapper software (PE Applied Biosystems). Instability was assigned to a marker if its fragment pattern displayed either additional peaks or the appearance of separated novel fragments when the profiles of normal and tumor tissue were compared. Instability observed using the six microsatellites markers for elucidating the LOH status of chromosomes 18q or 17p (TP53 locus) was also recorded, and a separate global microsatellite instability status was assigned with the 11 microsatellite markers.
In accordance with the consensus definitions of the US NCI, tumors were classified as exhibiting high microsatellite instability when 30% or more of the tested loci were unstable and as non-high microsatellite instability when less than 30% were unstable.
Statistical analyses
Clinicopathological variables were investigated for their possible association with the presence of micropapillary pattern in at least 5% of the tumor. Categorical variables were analyzed by Fisher’s exact tests of contingency tables, with odds ratio calculation as appropriate. Numerical variables were analyzed using the non-parametric Wilcoxon signed-rank test, which compares median differences. For all statistical tests, taking into account the exploratory character of our study, the P-values were considered as very strong evidence for P-values lesser than 0.001, a strong evidence for pPvalues between 0.001 and 0.01 and weak evidence for P-values between 0.01 and 0.05. Data were analyzed using the R package v.2.3.1 (R Development Core Team. 2006) supplied with the exactRankTest package. 26,27
Results
Clinicopathological findings
Table 1 summarizes the clinicopathological findings. Patients included 222 men and 157 women whose age ranged from 31 to 98 years (mean 68.7±12.0, median 70.0, IQ 17.0). Proximal location was seen in 137 tumors and 232 tumors were located in the distal colon. The location of 10 tumors was not specified.
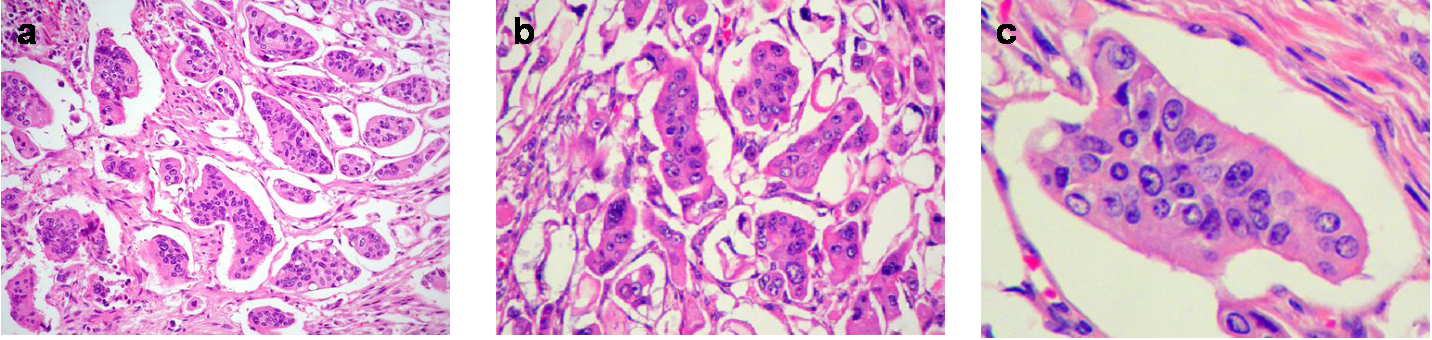
Microscopically, the micropapillary carcinoma pattern is composed of small irregular cell clusters surrounded by a thin lacunar space and separated by dense fibrous stroma (Figure 1a). Isolated infiltrating cells were also usually found. Tumor cells had a wide eosinophilic cytoplasm, a high-grade anaplasia (Figure 1b) and often showed a peculiar reverse cell polarity (Figure 1c). Immunohistochemistry for MUC1 (clone Ma695) in 85% of cases and MUC1 (EMA, clone E29) in 100% made this characteristic “inside-out” staining pattern in the micropapillary carcinoma component more evident (Figure 2a), whereas the rest of the tumor showed the classic luminal staining pattern (Figure 2b).
Figure 1 Micropapillary pattern. (a) Characteristic cell clusters surrounded by lacunar spaces and dense fibrous stroma (H&E, x 100). (b) MC cells displaying a high grade of anaplasia (H&E, x 400). (c) Tumor nest with reverse cell polarity (H&E, x 400).
Figure 2 (a) Immunohistochemistry for MUC-1 showing characteristic «inside-out» staining pattern in the MC component (x 400). (b) The rest of the tumor shows luminal staining pattern (MUC-1, x 400).
Micropapillary carcinoma involving at least 5% of the tumor volume was identified in 60 out of 379 colorectal carcinomas studied (16%) and was mainly located at the edge of the tumor. The proportion of this component varied, but in the majority of cases (95%) it represented less than 30% of the tumor. Out of 379 cases, 43 demonstrated between 5-10% micropapillary carcinoma pattern, 14 cases between 11-30% and only 3 cases more than 30%.
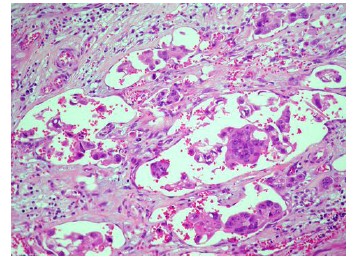
Comparison of clinicopathological features showed very strong evidence that pT (P=0.00088) and Astler-Coller stage (P=6.29 10-7) were higher in micropapillary colorectal carcinomas, grade (P=0.0018) was strongly related to the presence of micropapillary carcinoma, whereas mean age (P=0.0177) and tumor size (P=0.0429) were only weakly related. All tumors with a micropapillary carcinoma component displayed an infiltrative growth pattern and had more frequent nodal involvement (P= 8.49 10-9; Table 3) and peritoneal invasion (P=0.0012), as well as a higher percentage of the solid pattern (P=0.0003). There was a very strong evidence that micropapillary carcinomas presented perineural (P=0.0006), venous vessel (P=1.86 10-5) and thin-walled vessel invasion (P=7.47 10-12; Figure 3). The micropapillary carcinoma group more commonly had a high cytological grade (P=0.0031) and distant metastasis (P=0.0479). There was weak evidence of a higher percentage of ulcerated/stenosing tumors in the micropapillary carcinoma group (P=0.0236). No significant differences were found with respect to sex of the patient, location, presence of adenomas, tumor-infiltrating lymphocites, Crohn-like peritumoral reaction or the percentage of mucinous or cribriform patterns.
Figure 3 Thin-walled vessel invasion (H&E, x 200).
Table 3 Distribution micropapillary component vs metastasis
____________________________________________________________________
| % MC | Nodal metastasis (%) | Distant metastasis (%) |
| P-value | P-value | |
| 8.49 x 10-9 | 0.0479 | |
| <5% (n=319) | 127 (40) | 25 (8) |
| ≥5-10% (n=43) | 32 (74) | 7 (16) |
| >10-30% (n=14) | 13 (93) | 2 (14) |
| >30 (n=3) | 3 (100) | 1 (33) |
Abbreviation: MC, micropapillary component.
Bold values are statistical significance.
P-values reflect the different percentage of metastasis present in tumor with and whitout MC component.
Immunohistochemistry and molecular findings
KRT20 staining was positive in nearly all micropapillary carcinoma cases (97%) and in the majority (93%) of conventional adenocarcinomas. No statistical differences were found between the groups (P=1.00). Out of 30 conventional adenocarcinomas studied by KRT7, only 1 case of proximal location showed KRT7 expression, whereas 5 out of 30 micropapillary carcinoma cases were positive, 2 cases of proximal location and 3 cases of distal location. KRT7 expression was more frequent in micropapillary carcinoma than in conventional adenocarcinoma (17% vs. 3%), but this difference did not reach statistical significance (P=0.1967). KRT7 expression was often located at the peripheral areas of the tumor, corresponding to the transition areas from glandular to micropapillary carcinoma pattern. Finally, CEACAM5 expression was observed in all of the micropapillary carcinoma and conventional adenocarcinomas (Table 2).
The molecular findings are also summarized in Table 2. The difference in the expression of TP53 and MMR genes by immunohistochemistry did not reach significance. With weak evidence, a higher proportion of TP53 alterations (mutation and/or accumulation) was detected in the micropapillary carcinoma group (P=0.0343). However, no differences in the presence of KRAS or BRAF mutations were observed between carcinomas with and without micropapillary pattern, nor were differences found in the losses of 17p or 18q between the groups. There was a weak statistical evidence in the microsatellite instability statuses of these two groups of carcinomas using either the NCI microsatellite panel (P=0.0236) or the panel with 11 microsatellites (P=0.0123), whith high microsatellite instability classification being rare among micropapillary carcinoma cases.
Discussion
The histological micropapillary carcinoma profile described in other studies8-13 was characterized by small clusters of tumor cells with a clear lacunar space around the tumor nest (Figure 1a). The nuclei were moderately to highly pleomorphic, and the cytoplasm was abundant and showed eosinophilia (Figure 1B). The micropapillary focus was often associated with single cell infiltration. Many tumor nests had a peculiar reverse glandular polarity (Figure 1c), which was also confirmed immunohistochemically by the membranous staining pattern of MUC1 (clone MA695), towards the stromal pole only, in the tumor cell clusters (Figure 2a).
Micropapillary pattern has overlapping morphological features with the histological appearance of the tumor budding in the front of invasion of many colorectal carcinomas. However the later, which reflects the epithelium-mesenchymal transition at the edge of invasion, lacks the characteristic “inside-out” staining pattern for MUC1 observed in the micropapillary carcinoma.28
MUC1, a glycoprotein typically located in the apical cell surface of normal glandular epithelium, is thought to have an important role in lumen formation and generally to inhibit interaction between the cell and the stroma.8,14 Therefore, MUC1 interference with intercellular adherence is a key factor in the luminal formation of normal glands and probably also in the detachment of cells from the stroma. In invasive micropapillary carcinoma, the apical surface of inverted tumor cells faces the stroma and produces MUC-1, which directly interacts with the stromal cells. This process could have an important role in determining the characteristic morphological features of micropapillary carcinoma (lacunar space) and its invasive potential and aggressive behavior. 8,14,15
The aggressive profile of tumors with the micropapillary carcinoma component was highlighted in a study carried out by Haupt et al10 These authors found a major frequency of lymph node metastasis (P=0.001), lymphovascular invasion (P<0.050) and distant metastasis (P=0.311). Kim et al9 found an association between the presence of the micropapillary carcinoma component and lymphovascular invasion (P=0.011), nodal metastasis (P<0.001), major number of lymph nodes affected (P=0.006), advanced stage (P<0.001) and major frequency of distant metastasis (P<0.001). In agreement with these two reports,9,10 we found a significantly greater proportion of cases with lymphovascular invasion and a higher number of affected lymph nodes. Furthermore, we also demonstrated a highly significant association between the presence of micropapillary carcinoma and the extent of invasion, with most micropapillary carcinoma cases (92%) classified as either T3 or T4. The grade and overall stage were also significantly higher in themicropapillary carcinoma group. In addition, vascular (thin-walled and venous vessel) and perineural invasion were significantly more frequent in carcinomas with the micropapillary carcinoma component.
The incidence of micropapillary carcinoma in the literature ranges from 9 to 19%, and the proportion of the micropapillary carcinoma component with respect the whole tumor is also variable but usually less than 30% of tumor volume and mainly located at the periphery of the tumor.9,10
Here, we present an analysis of a series of 379 colorectal carcinomas, including 60 cases with at least 5% micropapillary component (16%). Most of our micropapillary carcinoma cases (95%) contained between 5 and 30% of the micropapillary carcinoma component, and no tumors were pure micropapillary carcinoma. These proportions are very similar to those obtained by Kim et al.9 and Haupt et al.10
The low proportion of micropapillary carcinoma component in the global tumor volume and its prognostic relevance makes the number of sections studied crucial to keep in mind. An increase in the number of sections routinely studied, especially sampling the tumor infiltrating edge, can likely result in a higher incidence of detection of the colorectal micropapillary carcinoma variant. In colorectal biopsies, as in the surgical specimens, it is very important to always report the presence of micropapillary carcinoma, however, small its proportion in the tumor, because the bad prognosis does not depend on this ratio, but rather on the sole presence of this component (Table 3).
In polypectomy specimens having adenoma with malignant transformation, the finding of a micropapillary component should also be emphasized as an indicator of more aggressive potential and a high probability of regional lymph node metastasis. In these situations, we agree with Sonoo et al., who recommend surgery as a more suitable procedure.29
With regard to the immunohistochemistry profile, Kim et al.9 do not report significant differences between colorectal carcinomas with or without a micropapillary carcinoma component. Nearly all cases of both micropapillary carcinoma and conventional adenocarcinoma showed immunoreactivity for KRT20 (95-100%), whereas KRT7 staining was usually negative (80-95%). In our immunohistochemistry study, a remarkable incidence of KRT7 expression (17%) was observed in the micropapillar carcinoma component, even when the subset of cases studied was slightly unbalanced to distal location. This finding could be significant when studying metastatic adenocarcinomas of uncertain origin with a micropapillary pattern. KRT20 expression is in agreement with colorectal carcinoma, ruling out other origins such as breast, lung or ovarian serous carcinoma.4,30,31 KRT7 immunoreactivity does not rule out possible colorectal origin. The expression of both KRT7 and KRT20 can suggest an origin in the pancreas or urothelium but, again, do not exclude colorectal carcinoma.
High instability in colorectal carcinoma has been associated with a good prognosis and typically is associated with an expansive growth pattern and absence of TP53 alterations. In our series, 58 out of 60 cases of micropapillary carcinoma had an infiltrative growth pattern and a higher proportion of TP53 alterations. The observed differences in the presence of TP53 mutations analyzed separately did not reach significance, although significance may be obtained with more extensive study. On the other hand, TP53 accumulations clearly showed no differences between carcinomas with and without a micropapillary pattern, but it is well known that there is not always a direct association between TP53 accumulation and the presence of TP53 gene mutations. Mutations that generate stop codons or frameshifts result in the absence of detectable protein expression, whereas TP53 accumulation can also be due to alterations in other proteins involved in the TP53 pathway. 32,33
When comparing the molecular profile of micropapillary carcinoma and conventional colorectal adenocarcinomas, although it did not reach a strong statistical evidence, microsatellite instability status was significantly different by direct assessment with two microsatellite panels (NCI or NCI+17p+18q), and significant differences were also detected when considering, as a whole, MMR loss and/or microsatellite instability status (RER phenotype). These data constitute a relevant finding of our molecular study because they demonstrate an association between the presence of the microsatellite carcinoma phenotype and the classical chromosomal instability pathogenic pathway and, consequently, a worse prognosis.
In conclusion, colorectal micropapillary carcinoma appears to be more aggressive than conventional colorectal adenocarcinoma, regardless of its proportion. Thus, we must highlight the importance of carrying out exhaustive research into the presence of this component in the pathological study of colorectal cancer.
Preliminary results from this article were published as a poster presentation at the 99thAnnual Meeting of the United States and Canadian Academy of Pathology (Washington, DC, USA, March 2010).
Acknowledgments
We thank Nuria Arraiza from Histopat for the execution of the immunohistochemical stains. We also thank Ana Mª Guardiola from Histopat and Eva Torija from Biopat for their secretarial assistance in data collection.
Disclosure/conflict of interest
The authors declare no conflict of interest.
References
- Siriangkul S, Tavasoli FA. Invasive micropapillary carcinoma of the breast. Mod Pathol 1993; 6:660-662.
- Amin MB, Ro YJ, el-Sharkawy T et al. Micropapillary variant of transitional cell carcinoma of the urinary bladder. Histologic pattern resembling ovarian papillary carcinoma. Am J Surg Pathol 1994;18:1224-1232.
- Paterakos M, Watkin WG, Edgerton SM, Moore DH, Thor AD. Invasive micropapillary carcinoma of the breast: a prognostic study. Hum Pathol 1999;30:1459-1463.
- Nassar H, Wallis T, Andea A, Dey J, Adsay V, Visscher D. Clinicopathologic analysis of invasive micropapillary differentiation in breast carcinoma. Mod Pathol. 2001;14(9):836-841.
- Amin MB, Tamboli P, Merchant SH et al. Micropapillary component in lung adenocarcinoma: a distinctive histologic feature with possible prognostic significance. Am J Surg Pathol 2002;26:258-364.
- Zekioglu O, Erhan Y, Ciris M, Bayramoglu H, Ozdemir N. Invasive micropapillary carcinoma of the breast: high incidence of lymph node metastasis with extranodal extension and its immunohistochemical profile compared with invasive ductal carcinoma. Histopathology 2002;44:18-23.
- Nassar H. Carcinomas with micropapillary morphology: clinical significance and current concepts. Adv Anat Pathol. 2004;11(6):297-303. Review.
- Sakamoto K, Watanabe M, De la Cruz C, et al. Primary invasive micropapillary carcinoma of the colon. Histopathology 2005; 47.479-484.
- Kim MJ, Hong SM, Jang SJ, et al. Invasive colorectal micropapillary carcinoma: an aggressive variant of adenocarcinoma. Hum Pathol 2006;37:809-815.
- Haupt B, Ro JY, Schwartz MR, Shen SS. Colorectal adenocarcinoma with micropapillary pattern and its association with lymph node metastasis. Mod Pathol 2007;20:729-733.
- Wen P, Xu Y, Frankel WL, Shen R. Invasive micropapillary carcinoma of the sigmoid colon: distinct morphology and aggressive behavior. Int J Clin Exp Pathol. 2008;1(5):457-460.
- Kuroda N, Oonishi K, Ohara M et al. Invasive micropapillary carcinoma of the colon: an immunohistochemical study. Med Mol Morphol 2007;40:226-230.
- Xu F, Xu J, Lou Z et al. Micropapillary component in colorectal carcinoma is associated with lymph node metastases in T1 and T2 stages and decreased survival time in TNM stages I and II. Am J Surg Pathol 2009;33:1287-1292.
- Nassar H, Pansare V, Zhang H, et al. Pathogenesis of invasive micropapillary carcinoma: role of MUC1 glycoprotein. Mod Pathol. 2004;17(9):1045-1050.
- Li YS, Kaneko M, Sakamoto DG, Takeshima Y, Inai K. The reversed apical pattern of MUC1 expression is characteristics of invasive micropapillary carcinoma of the breast. Breast Cancer. 2006;13(1):58-63.
- Hamilton SR, Aaltonen LA, eds. World Health Organization Classification of Tumors. Pathology and Genetics of Tumours of the Digestive System. Lyon; IARC Press; 2000:103-143.
- Sobin LH, Wittekind Ch, eds. TNM Classification of Malignant Tumors, 6th edition. New York: Wilsey-Liss;2002.
- Compton CC. Colorectal carcinoma: Diagnostic, prognostic, and molecular features. Mod Pathol 2003;16:376-388.
- Carethers JM, Smith EJ, Behling CA, et al. Use of 5-fluorouracil and survival in patients with microsatellite-unstable colorectal cancer. Gastroenterology. 2004;126:394-401.
- Valentini AM, Armentano R, Pirrelli M, Gentile M, Caruso M L. Immunohistochemical Mismatch Repair Proteins Expression in Colorectal Cancer. Appl Immunohistochem Mol Morphol 2006;14:42–45
- Shibata DK, Arnheim N, Martin WJ. Detection of human papillomavirus in paraffin-embedded tissue using the polymerase chain reaction. J Exp Med 1988; 167:225-230.
- Bauer HM, Ting Y, Greer CE, et al. Genital human papillomavirus infection in female university students as determined by a PCR-based method. JAMA. 1991;265:472-477.
- Boland CR, Thibodeau SN, Hamilton SR et al (1998) A National Cancer Institute Workshop on Microsatellite instability for cancer detection and familial predisposition: Development of international criteria for the determinationof microsatellite instability in colorectal cancer. Cancer Res 58: 5248-5257.
- Jen J, Kim H, Piantadosi S et al (1994) Allelic loss of chromosome 18q and prognosis in colorectal cancer. N Engl J Med 331: 213-221.
- Jones MH, Nakamura Y (1992) Detection of loss of heterozygosity at the human TP53 locus using a dinucleotide repeat polymorphism. Genes Chromosomes and Cancer 5: 89-90.
- R Development Core Team (2006). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, URL http://www.R-project.org.
- Torsten Hothorn and Kurt Hornik (2006). exactRankTests: Exact Distributions for Rank and Permutation Tests. R package version 0.8-17.
- Karamitopoulou E, Lugli A, Panayiotides I et al. Systematic assessment of protein phenotypes characterizing high-grade tumour budding in mismatch repair-proficient colorectal cancer Histopathology 2010;57: 233-243.
- Sonoo H, Kameyama M, Inatugi N, Nonomura A, Enomoto Y. Pedunculate Polyp of Early Sigmoid Colon Cancer with Invasive Micropapillary Carcinoma. Jpn J Clin Oncol 2009;39: 523-527.
- Tot T. Cytokeratins 20 and 7 as biomarkers:usefulness in discriminating primary from metastatic adenocaarcinoma. Eur J Cancer 2002;38:758-763.
- Logani S, Oliva E, Arnell PM, Amin MB, Young RH. Use of novel immunohistochemical markers expressed in colonic adenocarcinoma to distinguish primary ovarian tumors from metastatic colorectal carcinoma. Mod Pathol. 2005;18:19-25.
- Colomer A, Erill N, Verdú M, et al. The lack of p53 nuclear immunostaining is not indicative of absence of TP53 gene mutations in colorectal adenocarcinomas. Appl Immunohistochem Mol Morphol 2003;11:130-137.
- Greenblatt MS, Bennett WP, Hollstein M, Harris CC. Mutations in the p53 tumor suppessor gene: clues to cancer etiology and molecular pathogenesis. Cancer Res. 1994;54:4855-4878.